In this project, we looked for recent duplicated genes from Streptococcus mutans and Streptococcus sanguinis. Gene duplication event is considered recent if the duplication happens after the speciation event (See Figure 1). In this case, searching all proteins in an organism against proteins in non-redundant (NR) database using BLAST will identify the recently duplicated genes. The top hit will be against itself but if the second best hit comes from the same organism, we can consider this gene to be recently duplicated except in the case of gene loss for non-orthologous gene displacement1. Therefore, results shown here represents preliminary results that may need further analysis to be sure that the gene found is indeed recently duplicated.
The criteria for searching recent duplicated gene in Streptococcus mutans and Streptococcus sanguinis are as follows:
- Top-two hit organisms both belong to one of S. mutans and S. sanguinis
- Percentage of hit coverage for the length of protein must be greater than 80% and cutoff hit E-value less than 10-5.
From this search, S. mutans and S. sanguinis are found to have 16 and 29 putative duplicated genes in their genomes respectively. If the potentially duplicated gene pairs possess similar functional definition, we are more confident to say that they might be resulting from a gene duplication event. Bidirectional second best hit pairs for S. mutans and S. sanguinis are shown in the tables 1 and 2 below respectively.
Figure 1. Concept of recent gene duplication
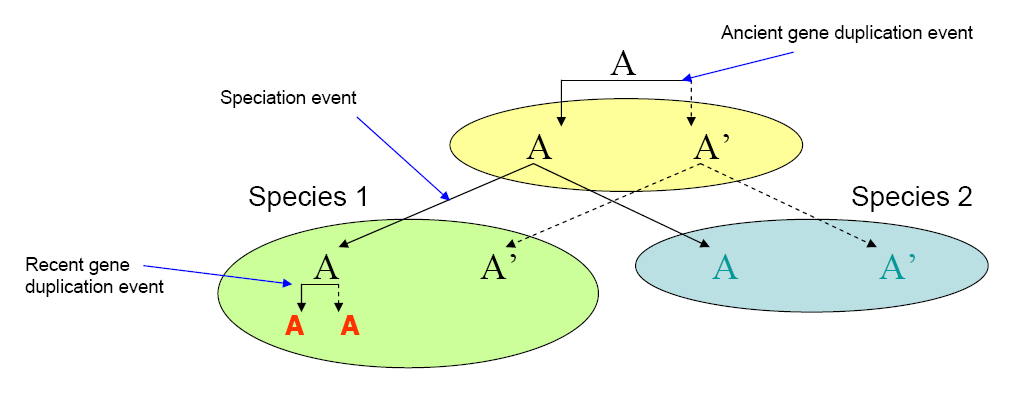
Table 1: The list of recent duplicated genes in Streptococcus mutans
| Locus Tag | Description | Hit Locus Tag | Hit Description | Hit Coverage | Hit E-value |
| SMU.875c | putative transposase, IS150-like | SMU.1370c | putative transposase, IS150-like | 0.980(201/205) | 1.00E-115 |
| SMU.1370c | putative transposase, IS150-like | SMU.875c | putative transposase, IS150-like | 0.980(201/205) | 1.00E-115 |
| SMU.436c | putative transposase, ISSmu1 | SMU.565c | putative transposase, ISSmu1 | 0.996(277/278) | 1.00E-157 |
| SMU.565c | putative transposase, ISSmu1 | SMU.436c | putative transposase, ISSmu1 | 0.996(277/278) | 1.00E-157 |
| SMU.767 | putative transposase, ISSmu1 | SMU.565c | putative transposase, ISSmu1 | 0.996(277/278) | 1.00E-157 |
| SMU.286 | putative ABC transporter, ATP-binding protein ComA | SMU.1881c | putative ABC transporter, ATP-binding protein | 0.811(619/763) | 0 |
| SMU.1881c | putative ABC transporter, ATP-binding protein | SMU.286 | putative ABC transporter, ATP-binding protein ComA | 0.834(634/760) | 0 |
| SMU.1347c | conserved hypothetical protein; possible permease | SMU.1347c. Best hit is SMU.1365c | conserved hypothetical protein; possible permease | 0.985(768/780) | 0 |
| SMU.1365c | hypothetical protein; possible permease | SMU.1347c | conserved hypothetical protein; possible permease | 0.985(768/780) | 0 |
| SMU.1021 | putative citrate lyase, alfa subunit | not found in smut gbk file, x-ray data | Putative Alfa Subunit Of Citrate Lyase | 0.960(498/519) | 0 |
| SMU.112c | putative transcriptional regulator | SMU.112c. Best hit gi is 93277277, a RpiR-like transcription factor. | putative transcriptional regulator | 0.884(220/249) | 1.00E-115 |
| SMU.1407c | putative transposase, ISSmu1 | SMU.565c | putative transposase, ISSmu1 | 0.996(277/278) | 1.00E-157 |
| SMU.1836 | 3-deoxy-7-phosphoheptulonate synthase | SMU.1837 | 3-deoxy-7-phosphoheptulonate synthase | 0.875(300/343) | 1.00E-156 |
| SMU.1893c | putative transposase, ISSmu1 | SMU.565c | putative transposase, ISSmu1 | 0.996(277/278) | 1.00E-157 |
| SMU.1954 | chaperonin GroEL | not found in smut gbk file, OME175 strain | GroEL | 0.930(489/526) | 0 |
| SMU.590c | putative transposase, fragment | SMU.1024c | putative transposase fragment | 0.943(50/53) | 7.00E-20 |
Table 2: The list of recent duplicated genes in Streptococcus sanguinis
| Locus Tag | Description | Hit Locus Tag | Hit Description | Hit Coverage | Hit E-value |
| SSA_0134 | Membrane carboxypeptidase (penicillin-binding protein), putative | SSA_0175 | Penicillin-binding protein 1B, putative | 0.935(701/750) | 0 |
| SSA_0175 | Penicillin-binding protein 1B, putative | SSA_0134 | Membrane carboxypeptidase (penicillin-binding protein), putative | 0.872(701/804) | 0 |
| SSA_1362 | ORFB, transposon ISSsa2 | SSA_0266 | ORFB, transposon ISSsa1 | 0.995(201/202) | 1.00E-111 |
| SSA_0266 | ORFB, transposon ISSsa1 | SSA_1362 | ORFB, transposon ISSsa2 | 0.995(201/202) | 1.00E-111 |
| SSA_1305 | hypothetical protein | SSA_1110 | hypothetical protein SSA_1110 | 0.870(160/184) | 8.00E-79 |
| SSA_1110 | hypothetical protein | SSA_1305 | hypothetical protein SSA_1305 | 0.870(160/184) | 8.00E-79 |
| SSA_1285 | hypothetical protein | SSA_1289 | hypothetical protein SSA_1289 | 0.975(154/158) | 5.00E-82 |
| SSA_1289 | hypothetical protein | SSA_1285 | hypothetical protein SSA_1285 | 0.975(154/158) | 5.00E-82 |
| SSA_1681 | ABC-type bacitracin resistance protein A, ATPase component, putative | SSA_1660 | ABC-type antimicrobial peptide transport system, ATPase component, putative | 0.992(251/253) | 1.00E-135 |
| SSA_1660 | ABC-type antimicrobial peptide transport system, ATPase component, putative | SSA_1681 | ABC-type bacitracin resistance protein A, ATPase component, putative | 0.969(251/259) | 1.00E-135 |
| SSA_1757 | hypothetical protein | SSA_1758 | hypothetical protein SSA_1758 | 0.842(170/202) | 4.00E-80 |
| SSA_1758 | hypothetical protein | SSA_1757 | hypothetical protein SSA_1757 | 0.842(170/202) | 4.00E-80 |
| SSA_0148 | Sugar ABC transporter, ATP-binding protein, putative | SSA_2040 | ABC transporter ATP-binding protein-multiple sugar transport, putative | 0.960(361/376) | 0 |
| SSA_0394 | hypothetical protein | SSA_0599 | hypothetical protein SSA_0599 | 0.849(73/86) | 8.00E-32 |
| SSA_0407 | ABC-type multidrug transport system (3-component subtilin immunity exporter), ATPase component, putative | SSA_0412 | ABC-type multidrug transport system (3-component subtilin immunity exporter), ATPase component, putative | 0.828(250/302) | 1.00E-112 |
| SSA_0432 | Formate--tetrahydrofolate ligase, putative | not found in ssan gbk file. From NCBI, this gi is SSA_0432. | FTHS1_STRSV Formate--tetrahydrofolate ligase 1 (Formyltetrahydrofolate synthetase 1) (FHS 1) (FTHFS 1) | 0.966(538/557) | 0 |
| SSA_0524 | Microcompartment protein, putative | SSA_0525 | Microcompartment protein, putative | 0.802(73/91) | 1.00E-29 |
| SSA_0555 | Conserved hypothetical cytosolic protein | SSA_0557 | hypothetical protein SSA_0557 | 0.942(129/137) | 5.00E-66 |
| SSA_0556 | hypothetical protein | SSA_0562 | hypothetical protein SSA_0562 | 0.932(96/103) | 9.00E-50 |
| SSA_0684 | Fibril-like structure subunit FibA, putative | SSA_1635 | hypothetical protein SSA_1635 | 0.958(660/689) | 1.00E-94 |
| SSA_0904 | CshA-like fibrillar surface protein A | SSA_0906 | CshA-like fibrillar surface protein C | 0.850(2268/2669) | 0 |
| SSA_0947 | hypothetical protein | SSA_0948 | hypothetical protein SSA_0948 | 0.832(164/197) | 9.00E-80 |
| SSA_0969 | hypothetical protein | SSA_0968 | hypothetical protein SSA_0968 | 0.858(103/120) | 5.00E-47 |
| SSA_1007 | ABC transporter ATP-binding protein-multiple sugar transport, putative | SSA_0148 | Sugar ABC transporter, ATP-binding protein, putative | 0.923(347/376) | 0 |
| SSA_1340 | Zn/Mn ABC-type porter lipoprotein, putative | SSA_1990 | Zn-porter lipoprotein, putative | 0.820(250/305) | 1.00E-121 |
| SSA_1369 | FmtA-like protein, putative | SSA_1366 | Alkaline D-stereospecific endopeptidase precursor, putative | 0.875(398/455) | 0 |
| SSA_1663 | Collagen-binding protein A | SSA_1666 | Collagen-binding surface protein, putative | 1.479(954/645) | 1.00E-170 |
| SSA_2016 | Phosphoglycerate mutase, putative | SSA_2015 | Phosphoglycerate mutase, putative | 0.828(192/232) | 8.00E-90 |
| SSA_2156 | Conserved uncharacterized protein | SSA_2155 | hypothetical protein SSA_2155 | 0.818(189/231) | 2.00E-87 |
Reference